For quotations, please use our online quotation form, and you may also contact us by
service@kendallscientific.com
+1-888.733.6849 (Toll-free)
+1-617.299.7367 (Int’l))
+1-888.733.6849
Our customer service representatives are available 24 hours, Monday through Friday to assist you.| Reactivity | Human Mouse Rat |
| Tested applications | WB IHC IF IP CHIP |
| Recommended Dilution | WB 1:500 - 1:2000 IHC 1:50 - 1:200 IF 1:20 - 1:50 IP 1:20 - 1:50 ChIP 1:20 - 1:100 |
| Calculated MW | 51kDa |
| Observed MW | Refer to figures |
| Immunogen | Recombinant protein of human IDH2 |
| Storage Buffer | Store at -20℃. Avoid freeze / thaw cycles. Buffer: PBS with 0.02% sodium azide, 50% glycerol, pH7.3. |
| Synonym | IDH; IDP; IDHM; IDPM; ICD-M; D2HGA2; mNADP-IDH; |
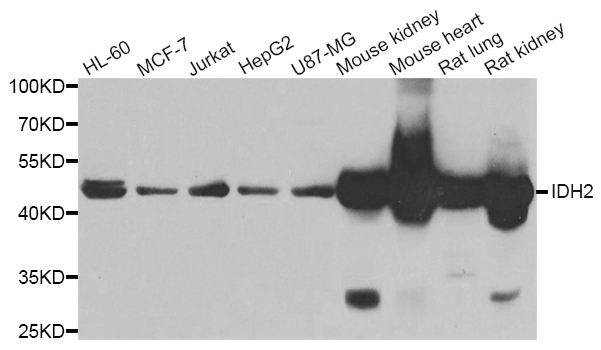
Western blot analysis of extracts of various cell lines, using IDH2 antibody.
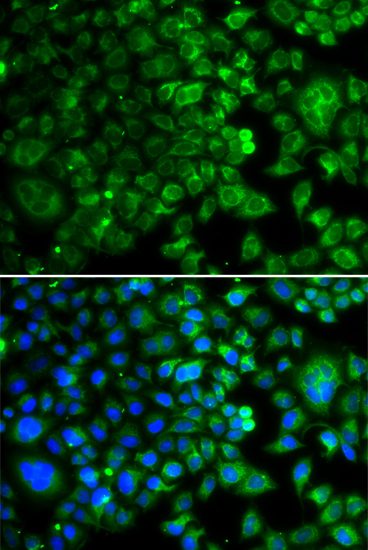
Immunofluorescence analysis of U20S cell using IDH2 antibody. Blue: DAPI for nuclear staining.

Chromatin immunoprecipitation analysis extracts of 293T cell line, using IDH2 rabbit polyclonal antibody and rabbit IgG. The amount of immunoprecipitated DNA was checked by quantitative PCR. Histogram was constructed by the ratios of the immunoprecipitated DNA to the input.
Isocitrate dehydrogenases catalyze the oxidative decarboxylation of isocitrate to 2-oxoglutarate. These enzymes belong to two distinct subclasses, one of which utilizes NAD(+) as the electron acceptor and the other NADP(+). Five isocitrate dehydrogenases have been reported: three NAD(+)-dependent isocitrate dehydrogenases, which localize to the mitochondrial matrix, and two NADP(+)-dependent isocitrate dehydrogenases, one of which is mitochondrial and the other predominantly cytosolic. Each NADP(+)-dependent isozyme is a homodimer. The protein encoded by this gene is the NADP(+)-dependent isocitrate dehydrogenase found in the mitochondria. It plays a role in intermediary metabolism and energy production. This protein may tightly associate or interact with the pyruvate dehydrogenase complex. Alternative splicing results in multiple transcript variants.
N/A